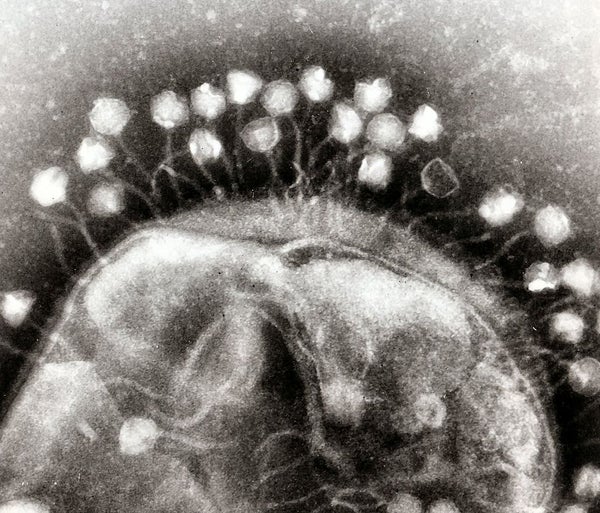
Viruses sense chemical signals left behind by their forebears so they can decide whether to kill or just to infect their hosts.
The discovery—in viruses that attack Bacillus bacteria—marks the first time that any type of viral communication system has ever been found. But researchers say that many other viruses could communicate with each other through their own molecular languages—perhaps even viruses that are responsible for human diseases. If that is the case, scientists might have found a new way to disrupt viral attacks.
The secret viral code was spotted by a team led by Rotem Sorek, a microbial geneticist at the Weizmann Institute of Science in Rehovot, Israel. Their findings are published in Nature on January 18.
On supporting science journalism
If you're enjoying this article, consider supporting our award-winning journalism by subscribing. By purchasing a subscription you are helping to ensure the future of impactful stories about the discoveries and ideas shaping our world today.
“This is going to be one of those transformative papers,” says microbiologist Martha Clokie, who studies viruses that infect bacteria (known as bacteriophages, or phages) at the University of Leicester, UK.
Infection signals
Sorek’s team was looking for evidence that a bacterium called Bacillus subtilis might alert other bacteria to phages. The researchers knew that bacteria speak to their brethren through secreting and sensing an array of chemicals. This phenomenon, called quorum sensing, allows the bacteria to adjust behaviours according to the numbers of other bacteria around. For instance, bacteria use quorum sensing to decide whether to divide or when to launch an infection.
Instead, the team found, to its surprise, that a viral invader ofBacillus bacteria—a phage called phi3T—makes a chemical that influences the behaviour of other viruses.
Some phages can infect cells in two different ways. Usually, they hijack host cells and multiply until the hosts burst and die. Sometimes, however, phages insert their own genetic material into a host’s genome, then lie dormant until a trigger causes them to reawaken and multiply later.
The newly discovered viral communication system alters the way phi3T infects.
The team first injected phi3T into a flask of Bacillus subtilis bacteria, and found that the virus tended to kill the bacteria. Then they filtered the contents of this flask to remove bacteria and viruses—but keeping small proteins—and fed this ‘conditioned medium’ to a fresh culture of bacteria and phages. That changed what the phage did: it was now more likely to slip its genome into the bacteria, rather than kill it. The team named the mysterious molecule that they suspected was involved ‘arbitrium’ (after the Latin word for decision) and set out to identify it.
‘Annoyingly good’
After a two-and-a-half year search, Sorek and graduate student Zohar Erez discovered that arbitrium was a short viral protein that seeps out of infected bacteria after death. When levels of arbitrium build up—after a large number of cells have died—phages stop killing off the remaining bacteria and retreat to lie dormant in bacterial genomes instead. Sorek, Erez and their colleagues identified two further phi3T proteins that measure levels of arbitrium and then influence the nature of subsequent infections.
“It does make a lot of sense,” says Peter Fineran, a microbial geneticist at the University of Otago in Dunedin, New Zealand. “If the phage is running out of hosts, it would try and limit its destruction, and sit quiet and wait for the host to re-establish growth.”
The new work is “annoyingly good”, says Clokie. “I’ve thought about doing those experiments to see if there’s something in the media.” She also expects other phage biologists will discover other communication systems. Sorek’s team found more than 100 different arbitrium-like systems, most of them in the genomes of other Bacillus viruses. “Phages broadcast in different frequencies. They speak in different languages and they can hear only the language that they speak,” he adds.
He even wonders whether viruses that infect more complex organisms, such as people, could talk to one another. HIV and herpes viruses can cause both active and latent infections, he notes. “If you had a molecule that could drive viruses into complete latency, it would be a good drug.”
This article is reproduced with permission and was first published on January 18, 2017.