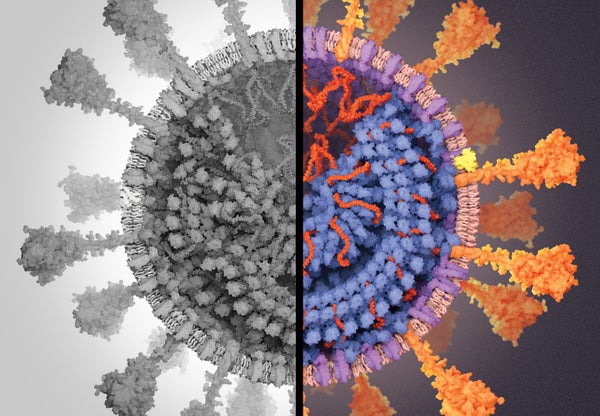
What does SARS-CoV-2, the virus that causes COVID-19, look like?
This simple question does not have a simple answer. SARS-CoV-2 is very small, and seeing it requires specialized scientific techniques. Electron microscopy (EM) can reveal its general size and shape. We can see that the virions are spherical or ellipsoidal, with “crowns” of spikes on their surfaces. Careful cryo-electron microscopy (cryo-EM) studies of many copies of the virion can reveal more precise measurements of the virus and its larger pieces. Those individual pieces can be studied separately from the virus, using cryo-EM, x-ray crystallography or NMR spectroscopy, resulting in atomic or near-atomic detail 3-D models.
However, flexible and disordered parts can evade even these techniques, leaving gray areas and ambiguity.
On supporting science journalism
If you're enjoying this article, consider supporting our award-winning journalism by subscribing. By purchasing a subscription you are helping to ensure the future of impactful stories about the discoveries and ideas shaping our world today.
Building a 3-D model of a complete virus like SARS-CoV-2 in molecular detail requires a mix of research, hypothesis and artistic license. Some structures are known, others are somewhat known, and others may be completely unknown.
When I was building the model shown in July’s issue of Scientific American, there were several places where I had to make best-guess decisions based on the evidence available. This model is not perfect; as scientific understanding of SARS-CoV-2 evolves, no doubt parts of it may need to be updated.
Many of the studies that this model is based on were done on SARS-CoV, the coronavirus that caused an outbreak known as SARS in 2003. SARS-CoV is closely related to SARS-CoV-2, and is structurally very similar. The differences in the diseases that they cause are probably the result of very small molecular features, which would barely be visible when looking at the virion as a whole. I decided at the outset to use SARS-CoV data as needed. The research on SARS-CoV-2 is still ongoing, and the very careful ultrastructural studies that have been done on SARS-CoV have yet to be done on SARS-CoV-2.
Here, I’ll walk through each component of the virion and review the evidence I found for its structure, and where I had to bridge gaps with hypotheses or artistic license.
OVERALL STRUCTURE
Under the electron microscope, SARS-CoV-2 virions look spherical or ellipsoidal. Scientists have measured diameters from 60 to 140 nanometers (nm). (This is about one thousandth the width of a human hair). However, the measurements available at the time of this model building were from negative-stain electron microscopy, which does not resolve detail as finely as cryo-EM.
For more precision measurements, I referenced a meticulously detailed cryo-EM study of SARS-CoV from 2006. There, researchers reported mean diameters of 82 to 94 nm, not including spikes. I ended up building my virion model to be spherical and 88 nm in diameter. This study also reported relative amounts of the structural proteins at the surface; each of these measurements are described, with the protein in question, below.
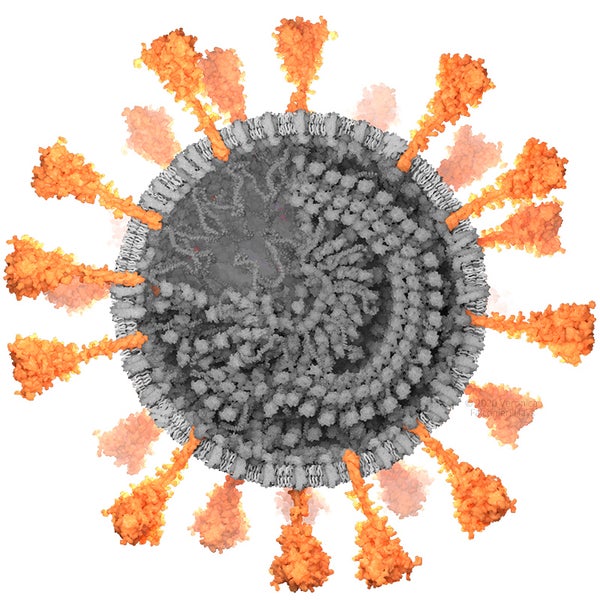
Spike (S) protein. Credit: Veronica Falconieri Hays
SPIKE (S) PROTEIN
The spike (S) protein sticks out from the viral surface and enables it to attach to and fuse with human cells. The top of the spike, including the attachment domain and part of the fusion machinery, had been mapped in 3-D by cryo-EM by two research groups (the Veesler Lab and McClellan Lab) by March 2020. However, the stem of the spike, the transmembrane domain and the tail inside the virion are not mapped. Scientists know that these regions exist, and what amino acids (protein building blocks) they include, but have not yet been able to observe their arrangement in 3-D space.
The inclusion of a stem is a key difference between my model and many SARS-CoV-2 visualizations. Most, including the iconic CDC image, use the 3-D data for the top of the spike but don’t show a stem, resulting in a shorter spike model.
At first, I modeled in a schematic stem, so the spike looked a bit like a rock candy lollipop. I matched it to the measured spike height and spacing from SARS-CoV, about 19 nm tall and 13–15 nm apart.
However, over on science Twitter, I had seen posts by Lorenzo Casalino, Zied Gaieb and Rommie Amaro, of the University of California, San Diego showing a molecular dynamics video of the spike and its attached sugar chains. They had built a complete spike model, including stem, transmembrane domain and tail, based on amino acid sequence similarity with known 3-D structures. This model was required for their molecular dynamics study (now in preprint) to learn more about how the spike behaves. They generously shared their model with me for inclusion in my visualization.
Notably, the Amaro lab model is 25 nm tall, 6 nm taller than I was expecting based on the measurements of SARS-CoV. I did not resolve this discrepancy, but my hypothesis is that, on actual virions, the spike stems bend and appear shorter under the electron microscope, and/or the flexibility of the very top of the spike blur its boundaries, which makes the height measurement somewhat ambiguous even by cryo-EM.
There is also a reported 9–12 nm height measurement of the SARS-CoV-2 spike based on a negative-stain EM image. However, negative-stain EM does not resolve detail as well as cryo-EM, which was used to make the 19 nm measurement. SARS-CoV-2’s spike also has a similar number of amino acids as SARS-CoV’s spike (1,273 versus 1,255), so it is very unlikely that SARS-CoV-2’s spike would be as small as these negative-stain based measurements suggest.
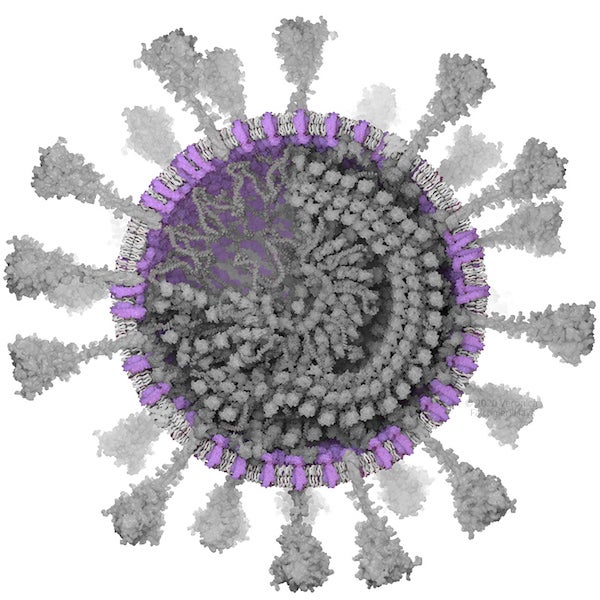
Membrane (M) protein. Credit: Veronica Falconieri Hays
MEMBRANE (M) PROTEIN
The membrane (M) protein is a small but plentiful protein embedded in the envelope of the virus, with a tail inside the virus that is thought to interact with the N protein (described below). Like the spike stem, the M protein has not been mapped in 3-D, nor has any similar protein. The SARS-CoV and SARS-CoV-2 M proteins are similar in size (221 and 222 amino acids, respectively), and based on the amino acid pattern, scientists hypothesize that a small part of M is exposed on the outside of the viral membrane, part of it is embedded in the membrane, and half is inside the virus. Based on this information, I assembled a model based on parts from two slightly similar proteins (Protein Data Bank entries 4NV4 and 5CTG as identified by SwissProt). The model for the intraviral domain had a long tail, but I could not confidently orient this and found it pointed out in odd directions, so I cut it off to avoid visual distraction or implication of a false structural feature.
The M proteins form pairs, and it is estimated that there are 16–25 M proteins per spike on the surface of the virus. I ended up modeling 10 M protein pairs (so 20 M proteins) per spike in my model. Some researchers hypothesize that the M proteins form a lattice within the envelope (interacting with an underlying lattice of N proteins; see below). I decided to use an icosahedral sphere to create a regular distribution of the M protein dimers to hint at this hypothesis.
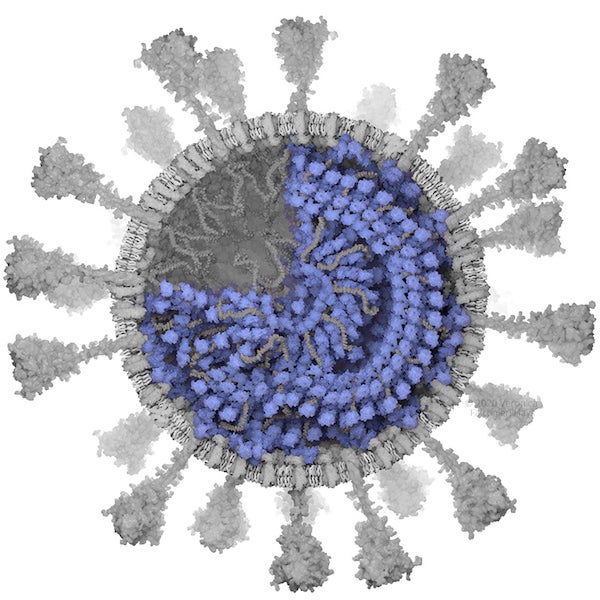
N protein. Credit: Veronica Falconieri Hays
N PROTEIN
The nucleoprotein (N protein) is packaged with the RNA genome inside the virion. It is thought to form a latticelike structure just beneath the envelope, and viral spikes can only fit between N proteins, preventing them from being spaced closer than 13–15 nm.
The N protein is made of two relatively rigid globular domains connected by a long disordered linker region. The structures of the two domains, the NTD and CTD, are known for SARS-CoV-2 and SARS-CoV, respectively, but exactly how they are oriented relative to each other is a bit of mystery. Based on the disorder of the linking domain, it could be highly variable.
The structure of the CTD was determined by x-ray crystallography, a technique that requires crystallizing purified copies of the protein. In this crystallization process, the CTD formed an interesting eight-piece structure, that, if stacked, forms a helical core. This is not definitive but highly suggestive that the viral RNA could wrap around this core. The N protein’s other half, the NTD, may then interact on the outside of the RNA, or, where it is close to the M protein and viral envelope, attach instead there. This would form the observed sub-envelope N protein lattice and would keep the entire RNA-N protein complex close to the membrane where possible.
The technical challenge of modeling hundreds of copies of N protein, each with two domains linked by disordered amino acid strings, was too great to be tackled while creating this model. I decided to place a lattice of NTDs beneath the viral spikes, build a core of helical CTDs for the RNA-N protein complex, and add NTDs both interacting with the RNA and scattered throughout the virion. I continued the spiral of the core into the center of the virus; this was my solution to packing in the extremely long RNA strand (more below), but in reality, the RNA and N protein may be more disordered in the center of the virion. The end result captures a few ideas of how the N protein is packed within, if not its full and dynamic complexity.
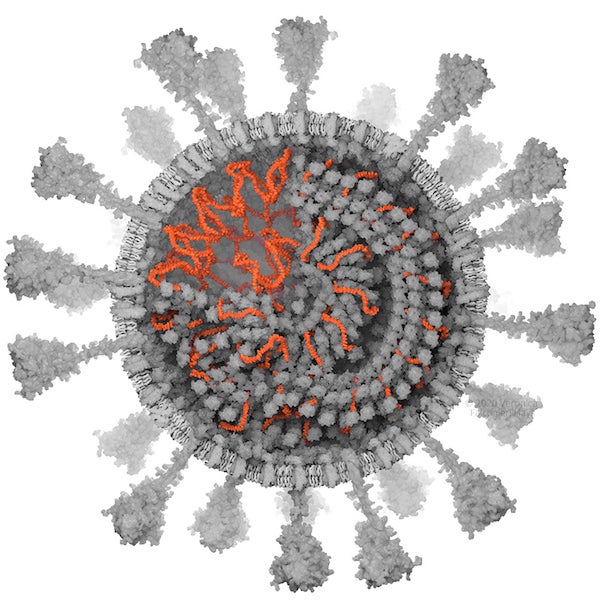
RNA. Credit: Veronica Falconieri Hays
RNA
At 29,903 RNA bases, SARS-CoV-2’s genome is very long compared to similar viruses. I wanted to make sure that my model of the RNA approximated the length of the genome. However, RNA structure can be complex; the bases in some regions can interact with others, forming loops and “hairpins” and resulting in very convoluted 3-D shapes. For this model, I made the assumption that the RNA was a stretched-out thread, neatly wrapped around an N protein core for its entire length. I found a research paper from 1980 that reported measurements of 4–4.8 RNA bases per nm, or about 3,000 to 3,750 nm for the half of the genome modeled into the virion cross section.
At first when I did this calculation, I was off by an order of 10. Fitting 300 nm RNA into the virion was a breeze! Upon review, Britt Glaunsinger, a virologist at the University of California, Berkeley, who was the project consultant, pointed out that there should be more RNA, and I revisited my calculations and caught my mistake. I needed to squeeze at least 3,000 nm into the 80 nm wide space within the virion cross section; this took a bit more 3-D finagling. Once I ran out of space near the periphery, I continued the spiral of the RNA and N protein into the center of the virion.
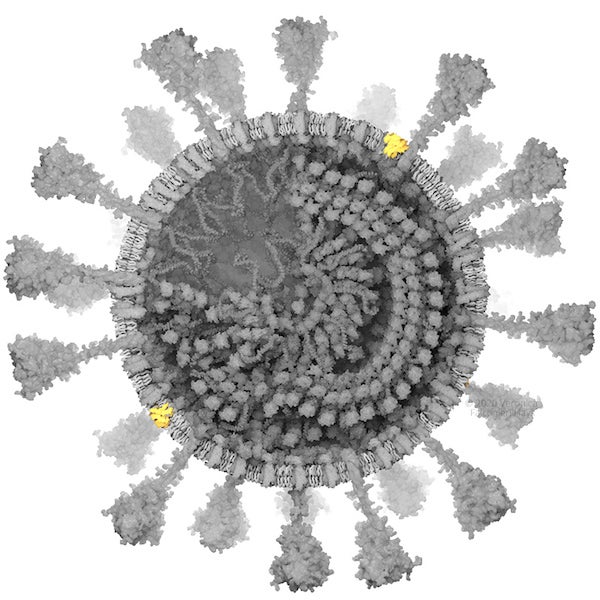
E protein. Credit: Veronica Falconieri Hays
E PROTEIN
The envelope (E) protein is a fivefold symmetric molecule that forms a pore in the viral membrane. Many copies are made during viral replication within the cell, but very few are incorporated into mature virions. Scientists have yet to map the SARS-CoV-2 E protein in 3-D, but there is an experimentally derived model of the SARS-CoV E protein, which is about 91 percent similar. I used that model here.
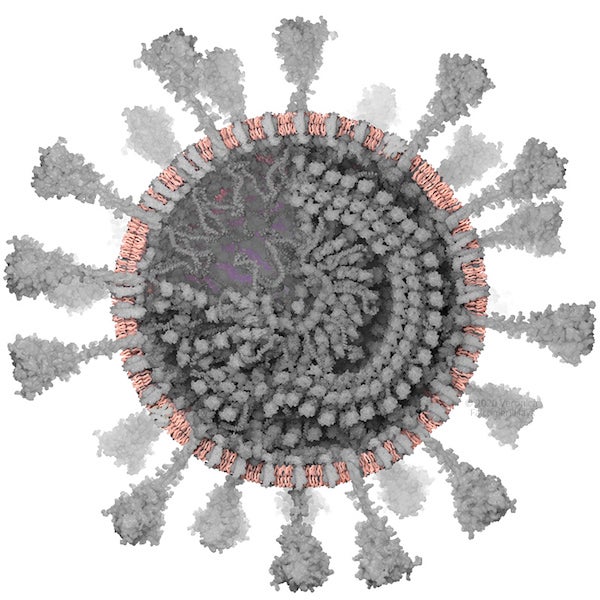
Envelope. Credit: Veronica Falconieri Hays
ENVELOPE
SARS-CoV-2 is enveloped in a lipid bilayer derived from organelle membranes within the host cell (specifically the endoplasmic reticulum and Golgi apparatus). I represented this with generic lipids: one head with two tails. With more time, this could have been more detailed. There are many different types of lipids, the proportions of which are specific to the membrane of origin.
BRINGING IT ALL TOGETHER
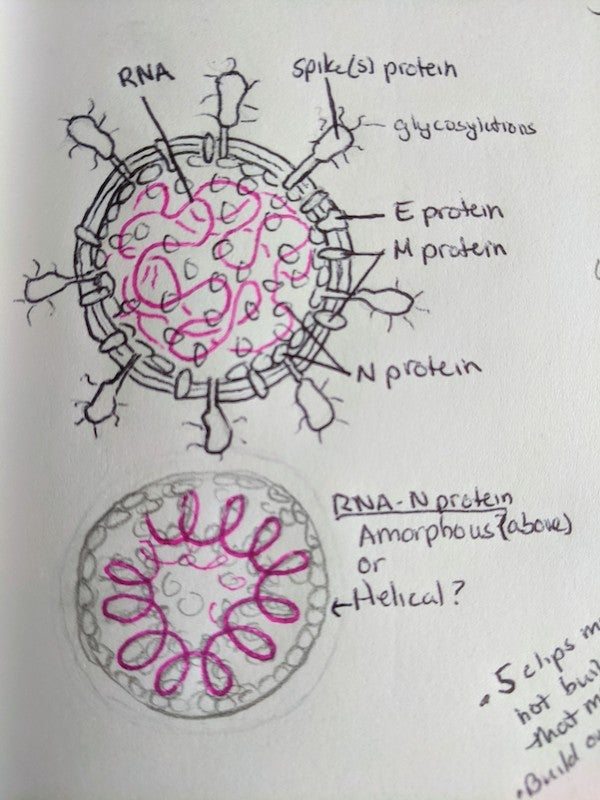
Virion sketch. Credit: Veronica Falconieri Hays
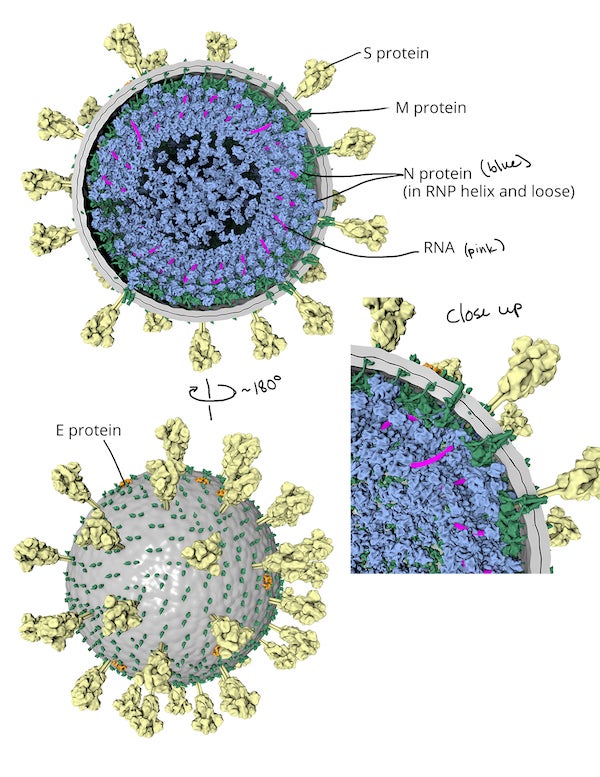
Virion draft, labeled. Credit: Veronica Falconieri Hays
ROUGH SKETCH AND INITIAL 3-D MODEL
After getting sign off on a quick hand-sketch of the virion to ensure all the necessary details were included, I started simultaneously researching and building the 3-D model in a 3-D modeling and animation program, Cinema4D. Within Cinema4D, I created an 88 nm sphere as a base, and then targeted copies of molecular models either on its surface or inside it. As my research progressed, I modified their distribution, and counted, measured and calculated as needed. I use the embedded Python Molecular Viewer (ePMV) plugin to import available 3-D molecular data directly.
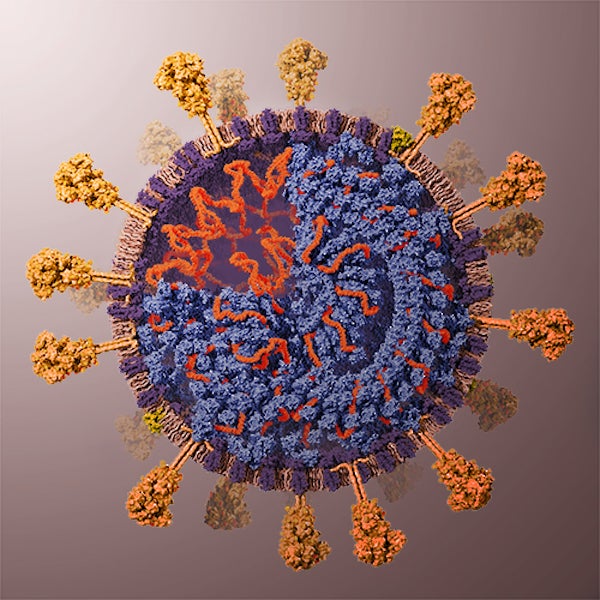
Render color guide. Credit: Veronica Falconieri Hays
INITIAL COLOR STUDY
I used a basic 2-D image of the resulting model to experiment with colors, and then used that palette as a starting point for creating my materials and setting up lighting in 3-D.
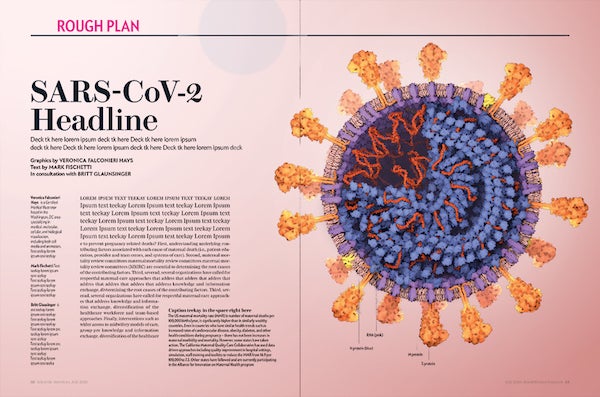
Color study 1. Credit: Veronica Falconieri Hays
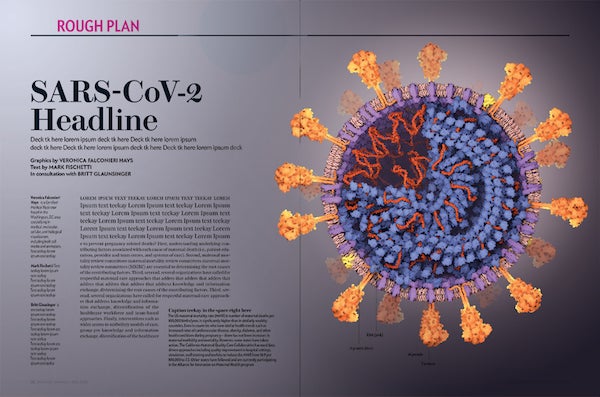
Color study 2. Credit: Veronica Falconieri Hays
3-D COLOR STUDIES
At first, I imagined a warm, pinkish background, as if looking closely into an impossibly well-lit nook of human tissue. However, I experimented in 2-D with a darker, cooler background and found I liked how it made the crown of spike proteins pop. Jen Christiansen, the art director, also liked this direction, so I refined the darker background version into the illustration found on the cover of the July 2020 issue of Scientific American.
Acknowledgments
I would like to acknowledge and thank my peers at the Association of Medical Illustrators (AMI) for sharing their research in an effort spearheaded by Michael Konomos. Thank you also to Nick Woolridge, David Goodsell, Melanie Connolly, Joel Dubin, Andy Lefton, Gloria Fuentes, and Jennifer Fairman for correspondence and visualizations that helped further my own understanding of SARS-CoV-2. Thank you to Scientific American’s Jen Christiansen for art direction, and for humoring the many deeply nerdy e-mails I sent her way during the making of this piece.