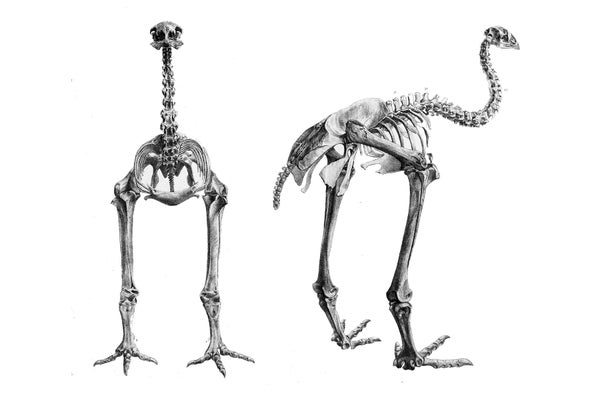
Scientists at Harvard University have assembled the first nearly complete genome of the little bush moa, a flightless bird that went extinct soon after Polynesians settled New Zealand in the late 13th century. The achievement moves the field of extinct genomes closer to the goal of “de-extinction”—bringing vanished species back to life by slipping the genome into the egg of a living species, “Jurassic Park”-like.
“De-extinction probability increases with every improvement in ancient DNA analysis,” said Stewart Brand, co-founder of the nonprofit conservation group Revive and Restore, which aims to resurrect vanished species including the passenger pigeon and the woolly mammoth, whose genomes have already been mostly pieced together.
For the moa, whose DNA was reconstructed from the toe bone of a museum specimen, that might require a little more genetic tinkering and a lot of egg: The 6-inch long, 1-pounder that emus lay might be just the ticket.
On supporting science journalism
If you're enjoying this article, consider supporting our award-winning journalism by subscribing. By purchasing a subscription you are helping to ensure the future of impactful stories about the discoveries and ideas shaping our world today.
The work on the little bush moa has yet to be published in a journal (the researchers posted a non-peer-reviewed paper on a public site), but colleagues in the small world of extinct genomes sang its praises. Morten Erik Allentoft of the Natural History Museum of Denmark, an expert on moa DNA and other extinct genomes, called it “a significant step forward.” Beth Shapiro of the University of California, Santa Cruz, who led a 2017 study reconstructing the genome of the passenger pigeon, called it “super cool” because it “gives us an extinct genome on an evolutionary branch where we hadn’t had any before.”
That the assembly of an extinct genome is being spread like scientific samizdat is not unusual in this field. Journals demand more from papers than “here it is,” said Ben Novak, a co-author of the passenger pigeon study. “The number that’s actually been done is possibly quadruple” the four or five extinct genomes formally reported, “but the results are just sitting in people’s labs.”
The nearly complete extinct genomes include two human relatives, Neanderthals and Denisovans, in addition to the woolly mammoth, and the passenger pigeon. The zebra-like quagga was the first extinct species to have its DNA sequenced, back in the genomic Stone Age of 1984, but it’s not up to modern standards.
Scientists are also close to reconstructing the genomes of the dodo, the flightless bird that went extinct from Mauritius, its only home, in the late 1600s; and the great auk, which lived in the North Atlantic before dying out in the mid-19th century. Last month, researchers in Australia unveiled the genome of the Tasmanian tiger, the last of which died in captivity in 1936.
In each case the steps were similar. Scientists collect tissue samples from museum specimens: the Museums Victoria in Melbourne, Australia, had great Tasmanian tigers, for instance, while the Royal Ontario Museum in Toronto had a nice toe bone from the little bush moa. They then extract DNA. It’s almost always as badly fragmented as a shattered wine goblet because “DNA decay begins within days of death,” said UCSC’s Shapiro.
Luckily, that’s not a problem. Today’s high-throughput genome sequencers actually work best on DNA measuring scores to hundreds of nucleotides—the iconic A’s, T’s, C’s, and G’sthat comprise DNA — long.
The tricky part is figuring out where the pieces belong on the genome: on which chromosomes and in what order. To do that, Harvard’s Alison Cloutier and the rest of the little bush moa team (which declined to talk about the work before its formal publication) took their 900 million nucleotides, scattered across millions of DNA pieces, and tried to match them to specific locations on the genome of the emu, a close relative of all nine moa species.
That enabled the scientists to get roughly 85 percent of the genome in the right place. “The other 15 percent is in their data but is difficult to organize using the emu genome,” said Novak. Turning tiny bits of DNA into a complete genome “has been extraordinarily difficult. The fact that they could get a genome from a little bush moa toe bone is a big deal, since now we might be able to use their data to do other extinct bird species.”
That’s because bird genomes, including the eight other (all extinct) moa species, have similar structures. That is, genes for particular traits tend to be on the same chromosome and arranged relative to other genes in a similar way. The more clues to how to organize the bits of genome that a sequencer spits out, the better.
For the passenger pigeon, for instance, Shapiro and her paleogenomics team used the genome of the band-tailed pigeon to figure out how to organize their short DNA sequences. She is trying to do something similar for the dodo, using the genome of the nicobar pigeon (the closest living species) as a template.
It’s “extremely difficult” to get genome organization right, said Charlie Feigin, a postdoctoral fellow at Princeton University who led the sequencing of the Tasmanian tiger genome. “You can look at closely related species for clues,” but with no guarantee of getting the extinct genome arranged correctly. “That structure does matter, but the extent to which it has to be perfect [for de-extinction to succeed] is debated.”
For the mammoth project, for instance, scientists are sequencing elephant chromosomes to get a better sense of how mammoth DNA is organized, said Harvard’s George Church, who is leading that project. They’re also hoping to improve on nature, genetically engineering resistance to the herpes virus into the mammoth genome (the better to keep it alive should de-extinction work). Some scientists believe that herpes infections helped kill off the mammoth. Church said his lab aims to announce progress on both fronts this year.
The best guess is that, if scientists resurrected an extinct species by putting its reassembled genome into the egg of a living species, it would likely not be a perfect replica of the original. A “de-extinct” passenger pigeon might eat what the original did but have different reproductive and social behaviors, for instance.
The putting-into-an-egg step turns out to be harder in birds than mammals. A reconstructed genome can be introduced into a mammalian egg with the cloning technique that produced Dolly the sheep. But that doesn’t work in birds—“at least so far,” said Brand. One hope is to get a workaround that recently succeeded in chickens, basically putting the genome into embryo cells that become eggs or sperm, to succeed in wild birds.
That’s “one of the things Revive and Restore is focused on now,” Brand said. “De-extinction is coming, gradually and certainly. It will eventually be seen as just another form of reintroduction,” like bringing “wolves back to Yellowstone Park [and] beavers back to Sweden and Scotland.”
Republished with permission from STAT. This article originally appeared on February 27, 2018