From Quanta Magazine (find original story here).
The world’s simplest known animal is so poorly understood that it doesn’t even have a common name. Formally called Trichoplax adhaerens for the way it adheres to glassware, the amorphous blob isn’t much to look at. At just a few millimeters across, the creature resembles a squashed sandwich in which the top layer protects, the bottom layer crawls, and the slimy stuffing sticks it all together. With no organs and just a handful of cell types, the most interesting thing about T. adhaerens might just be how stunningly boring it is.
“I was fascinated when I first heard about this thing because it has no real defined body,” said Michael Eitel, an evolutionary biologist at the Ludwig Maximilian University in Germany. “There’s no mouth, there’s no back, no nerve cells, nothing.”
On supporting science journalism
If you're enjoying this article, consider supporting our award-winning journalism by subscribing. By purchasing a subscription you are helping to ensure the future of impactful stories about the discoveries and ideas shaping our world today.
But after spending four years painstakingly reconstructing the blob’s genome, Eitel might know more about the organism than anyone else on the planet. In particular, he has looked closely enough at its genetic code to learn what visual inspections failed to reveal. The variety of creature that biologists have long called T. adhaerens is really at least two, and perhaps as many as a dozen, anatomically identical but genetically distinct “cryptic species” of animals. The discovery sets a precedent for taxonomy, the science of naming organisms, as the first time a new animal genus has been defined not by appearance, but by pure genetics.
The modern taxonomic system, little changed since Carl Linnaeus laid it out in the 1750s, attempts to chop the sprawling tree of life into seven tidy levels that grant every species a unique label. The two-part scientific name (such as Homo sapiens) represents the tail end of a branching path through this tree, starting from the thickest limbs, the kingdoms, and ending at the finest twigs, the genus (Homo) and then the species (sapiens). The path tells you everything there is to know about the organism’s relationship to other groups of creatures, at least in theory.
Ever since its discovery in the late 1800s, T. adhaerenshas been recognized as having a highly unusual body plan, and it has formally had the phylum of Placozoa (“flat animals”) to itself for almost half a century. Just one level more specific than kingdom, a phylum is a cavernous space to occupy alone: Our phylum, Chordata, overflows with more than 65,000 living species ranging from peacocks to whales to eels. Biologists have long suspected that Placozoa hid more diversity, and mitochondrial evidence strengthened that suspicion in 2004, when researchers found that short sequences from different individuals looked about as different as those of organisms from different families (one level more general than genus).
But that observation about the two Placozoa didn’t meet the accepted international standards for putting them in new taxonomic categories, which have historically been based on animals’ forms. “At the time we had just uncovered the genetic differences,” said Allen G. Collins, a co-author of the 2004 paper and a zoologist at the National Systematics Laboratory of the National Oceanic and Atmospheric Administration (NOAA). “Looking at the animals we had collected, it wasn’t discernible how they might differ morphologically.”
To finish what Collins started, Eitel and his colleagues decided to abandon the visual approach and search for defining characteristics in the placozoan genome itself.
They began by mapping out the phylum’s genetic territory with the same easy-to-sequence mitochondrial DNA Collins had used. By comparing data from this molecule, known as 16S, Eitel concluded that a particular variety of Placozoa from Hong Kong was the most distant relative of the standard strain, the genome of which had already been fully sequenced in 2008. If any group would qualify as a different species, this was the one.
He next needed to read, order and interpret the 80-odd million A, G, C and T nucleotide bases that make up the Hong Kong variant’s genome. Growing a few thousand placozoans, blending them to extract their nuclear DNA and converting the snippets of their genome into a digital format took a few weeks, but the hard work of shuffling those pieces into the right order and figuring out what each section does took four years of fiddling around with computer programs. When the team finally had a full genome ready for comparison, the payoff turned out to be worth the wait. “We expected to find differences, but when I first saw the results of our analyses, I was really overwhelmed,” Eitel said.
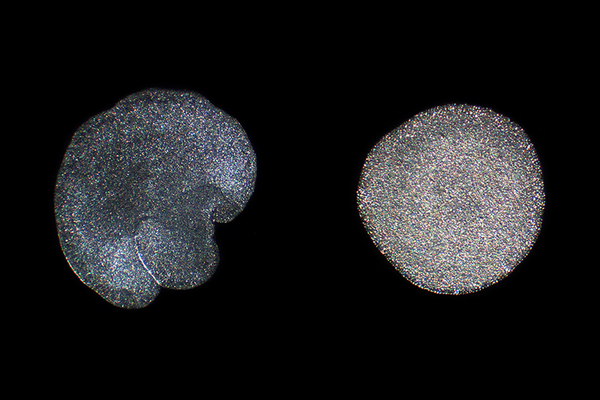
A quarter of the genes were in the wrong spot or written backward. Instructions for similar proteins were spelled nearly 30 percent differently on average, and in some cases as much as 80 percent. The Hong Kong variety was missing 4 percent of its distant cousin’s genes and had its own share of genes unique to itself. Overall, the Hong Kong placozoan genome was about as different from that of T. adhaerens as human DNA is from mouse DNA. “It was really striking,” Eitel said. “They look the same, and we look completely different from mice.”
So where do all those genetic changes manifest, if not in the animals’ flabby appearance?
“Even though the placozoan itself looks like a little ball of glue, it probably has cells that are doing some pretty sophisticated things,” said Holly Bik, a marine biologist at the University of California, Riverside, who studies tiny marine roundworms known as nematodes, which can also be cryptic. The Hong Kong Placozoa came from a brackish mangrove stream where large swings in temperature and salinity demand flexible body chemistry. “Physiologically, for organisms, that’s a pretty big thing to have to deal with. At the molecular level you need specific adaptations,” said Bik, who was not involved in the research.
By comparing the Placozoa variation with the average genetic differences between groups in other phyla, the German team concluded that the Hong Kong Placozoa qualified as not only a new species, but also a new genus. It might even have qualified as a new family or order in other areas of the animal tree, but to err on the conservative side, the team based their standard of genus variation on jellyfish, a genetically diverse phylum with relatively tidy divisions between levels.
All that remained was the naming. Taxonomic codes demand identifying characteristics, but don’t specify whether they should be visual or genetic, so the team picked out four genetic letters in the 16S mitochondrial genome that could uniquely differentiate the two lineages. Then, endorsed by peer review and PLOS Biology in late July, their work placed a new organism on our map of life.
The team gave their specimen the genus name Hoilungia, for a shapeshifting dragon king from Chinese mythology, and they named the species hongkongensis, for where it was collected. Similar genome-based classifications are common in the protist and bacterial worlds, and a relative handful of cryptic animal species have been named based on genetics. Namings (and renamings) that blend morphological characters with genetic ones, which recently re-classified a common houseplant, are also growing more common. But this was the first time genetic characters alone, unsupported by features like beak size or fin number, had been used to define an animal genus. “These people did the whole thing from the sexy molecular biology all the way to the proper naming,” said Susanne Renner, a botanist at the Ludwig Maximilian University. “It’s just great.”
The researchers hope their work will make it easier for future genetic character–based naming, which is less subject to biases from attention-grabbing visual features like antlers and fins that may not accurately reflect evolutionary distance between groups. “Someone had to be the first one to fight for the right to define new general species based on genomics, and we luckily got it published,” Eitel said.
Renner says this work is the latest step in an ongoing shift toward genetic taxonomy. “It took a long time to take off and now it’s taking off,” she said. She points out that in contrast with the pages of text that can go into a formal description of a species, specifying an organism with just four letters as the German team has done lends itself to snappy efficiency. “Linnaeus would be happy to do it. He was envisioning very brief and sharp diagnoses.”
As precise as genetic classification can be however, it will likely complement traditional ways of telling animals apart, not replace them. Observing visual features doesn’t require years of a lab’s time. Even for other cryptic animals like nematodes, which can’t be raised in captivity, genetic techniques may find limited use. “For me, working with a single nematode worm, there’s never going to be enough DNA isolated from an individual to use some of these technologies,” Bik said.
But for cryptic animals that researchers can cultivate, genetic sequencing may be the perfect spotlight for illuminating the shaded parts of their evolutionary tree. Eitel said he learned a lot from the process of analyzing the H. hongkongensis genome and predicts that sequencing the next variant—a project already underway—will take months, not years. “There will probably be dozens of new species popping up in the future,” he said. “And more to come, because we’re constantly sampling.”
Reprinted with permission from Quanta Magazine, an editorially independent publication of the Simons Foundation whose mission is to enhance public understanding of science by covering research developments and trends in mathematics and the physical and life sciences.