It began as an effort to understand how microbes fight viral infections. Within their chromosomes bacteria store snippets of DNA taken from the viruses they encounter. These fragments, which are tagged by a set of DNA segments called CRISPRs (clustered regulatory interspaced short palindromic repeats), serve as a record of past infections, and allow bacteria to become immune to future infections.
For Jennifer Doudna, an HHMI investigator and professor of chemistry as well as biochemistry and molecular biology at the University of California, Berkeley, the big question was, how did the system work? The answer lay with an enzyme named Cas9. Doudna and her team found that when armed with an RNA copy of one of the viral mug shots, the Cas9 enzyme could recognize and disable viruses that carried a matching sequence.
Once she understood this system, Doudna set out to harness it. By feeding the Cas9 enzyme a guide RNA of her choosing, Doudna found that she could edit target DNA much more easily and accurately than with existing methods. A Cas9-directed incision could inactivate a target gene. It could also provide an insertion site for new DNA, such as an altered version of the target gene.
The description of this CRISPR- Cas9 system—published in Science in 2012—launched a revolution in biology and biotechnology. In just seven years, CRISPR has become an essential research tool and the inspiration for scores of new start-ups. The technology has the potential to transform basic science, improve agricultural crops and cure genetic diseases. At the same time, it raises ethical questions about how to handle a technology that has the power to alter human evolution.
In 2018, Doudna and two of her colleagues, Professors Emmanuelle Charpentier at the Max Planck Institute for Infection Biology and Virginijus Siksnys at Vilnius University, were awarded the Kavli Prize in Nanoscience for their work on CRISPR-Cas9.
Now, Doudna outlines the next big questions that need to be addressed before CRISPR can reach its full potential.
What is CRISPR 2.0, and how do we develop it?
One big issue with CRISPR technology is how we can ensure the accuracy and the efficiency of genome editing, meaning the exact changes that are introduced into DNA. Right now, scientists can trigger targeted changes to DNA at a particular place in the genome, but we have a harder time ensuring the exact kind of change that gets introduced. A couple of things that are in the pipeline right now, not just in my own lab, but generally in the field: One is to develop what are called base-editing versions of CRISPR-Cas proteins. This means developing ways of using these programmable enzymes, not to cut DNA, but actually to trigger a chemical change to a particular DNA base in a sequence. With these base-editing molecules, we could reduce opportunities for the cell to make an undesired change. This is the kind of very specific manipulation of a DNA sequence that could, in principle, cure a disease-causing mutation in a cell.
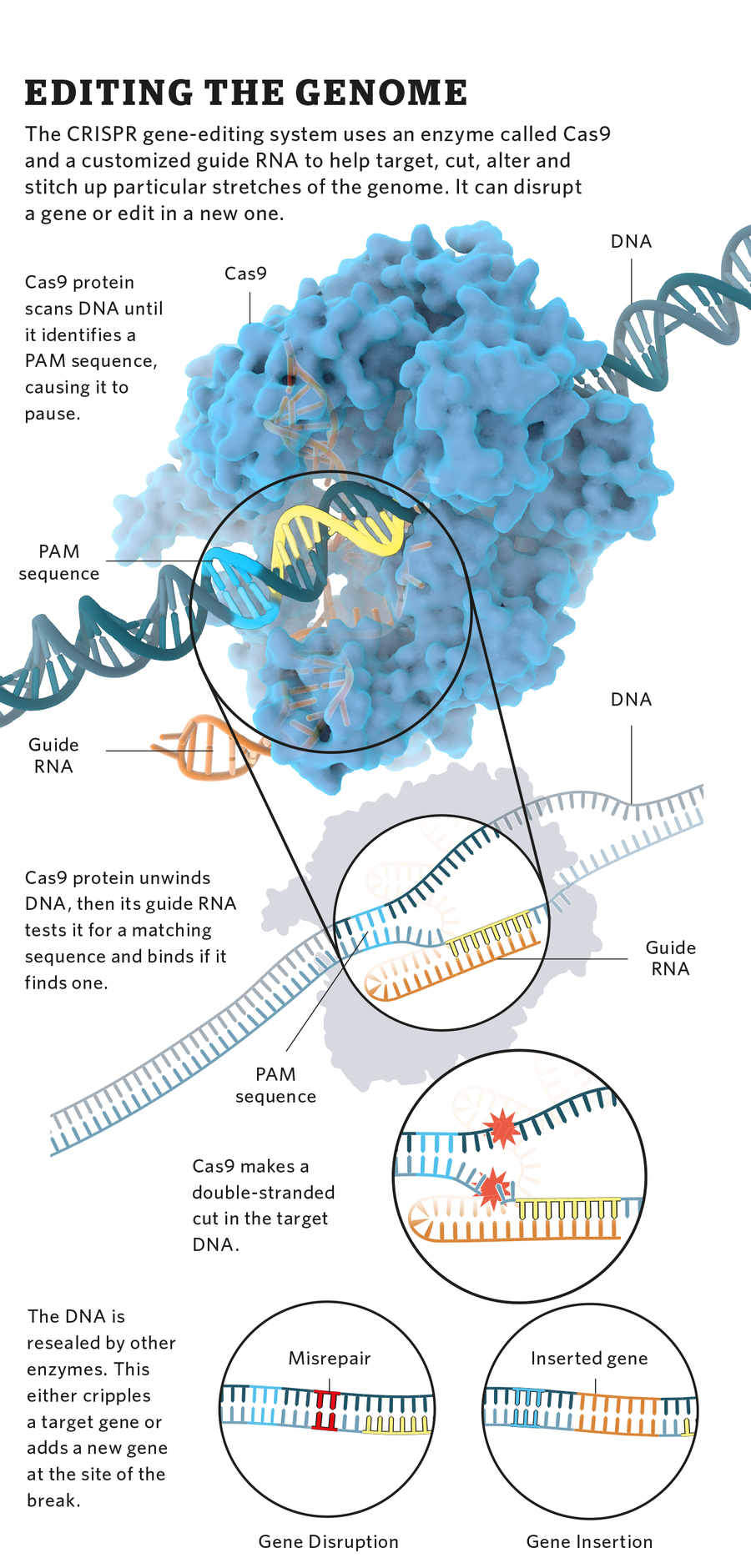
Illustration by Falconieri Visuals
Can we turn CRISPR-Cas9 against infectious disease?
There’s been a lot of interest in asking the question, at least in a research setting. Could you harness the adaptive immune functions of CRISPR-Cas systems for protecting other kinds of cells from viruses? I think that is not too likely, at least in its present form, because viruses have a remarkable ability to adapt and evolve resistance to targeting mechanisms, such as the one used by CRISPR-Cas. On the other hand, do I think that there may be ways to target bacteria that are infectious agents in people? Absolutely. One of the forefronts of the field is to explore how we could use CRISPR-Cas systems to target some of the bacteria that are harmful to people.
Can we turn to microbes to find alternative gene-editing tools?
One of the amazing questions in biology is, what are all of the microbes that populate our planet? Many scientists, including my colleague, Jillian Banfield here at University of California, Berkeley, are studying microbes in the environment by sequencing their DNA and piecing together information about their lifestyles, community partners and environmental niches. That’s something where I think a biochemist and structural biologist like myself can engage with experts in DNA metagenomics to try to understand the molecular pathways in these organisms. Some of these pathways provide a defense against viruses, like CRISPR systems do. CasX is a newer iteration of CRISPR-Cas that can be programmed to find and cut DNA just like Cas9. But it’s a lot smaller, and it has a completely different molecular shape, so it may be easier to deliver it into cells and ensure that it does the accurate editing necessary for clinical use.
How can we ensure gene editing benefits everyone?
Very soon, we will be faced with many opportunities for manipulating DNA—not only in individuals but also in the cells that can transmit changes to future generations. Given that, I’d like to see a lot more opportunities for public interaction with scientists and more opportunities to explore the broader implications of gene editing. How does it affect the inequalities that we see across society? How does it affect people’s decisions about reproduction and genetic disease? How do we even define genetic disease? What do we consider to be health versus disease? Those kinds of questions need to be openly debated.
To hear the complete podcast with Jennifer Doudna, watch the video below.
To learn more about brilliant work of Kavli Prize Laureates, visit The Kavli Prize. To explore more of the biggest questions in science, click here.



